Purpose: to make RFP from jelly fish in bacteria and to learn about steps in genetic engineering
Materials and procedure:
Lab 2a can be found in the Amgen lab manual part 2a
Lab 4a can be found in the Amgen lab manual part 2a
Lab 5a can be found in the Amgen lab manual part 5a
Lab 6a can be found in the Amgen lab manual part 6a
Experimental overview:
part 2a: we used restriction digest to verify the plasmid.
-cut plasmid with BamH1 and Hind III to remove RFP-ara from bacterial plasmid
part 4a: we used electrophoresis to verify digest
-to make sure that we had cut the plasmid correctly, we used electrophoresis. This is a process that allows you to run the plasmid in a gel with a DNA ladder.
part 5a: transformation of bacteria with recombinant plasmid
-the biggest part of the lab was in this step. We made the bacteria into the recombinant plasmid. We needed to use enzymes to cut the ligase and plasmid to paste the gene of interest.
part 6: purification of RFP using chromotography
-the last step was to use chromotography to separate the RFP from the rest. We did this by using a column with hydrophilic beads. This allowed the RFP to collect in the resin bed. We then were able to separate the RFP from the beads using a buffer.
Data:
2a before the lab:
1. RFP with pBAD and Ara-C with ori with Amp-R are the two fragments. The RFP and pBAD is 807 Base Pairs, but Ara-C, ori, and Amp-R are 4495 Base Pairs.
2. The components that are needed in the plasmid are Ara-C and the RFP gene.
3. To determine which bacteria will survive you need to use the selectable marker.
2a Stop and Think:
1. The purpose of setting up the tube without the enzymes is to use it as a control sample.
2. The enzymes might work best at 37*C because that is the way it works in the human body so they are comfortable.
2a Questions:
1. Ori = origin of replication; RFP = red fluorescent protein (gene of interest); Amp-R = selectable marker; Ara-C = binds to promoter
2. The restriction enzymes act as a sort of defense weapon.
3. Bacteria would retain certain enzymes to allow them to live. It is making it more difficult in medicine today because the bacteria have become resistant and it is harder to kill them.
4. All follow the same central dogma DNA to mRNA through RNA polymerase, and mRNA to a protein.
5. Put ampicillin and Kanamycin on a petri dish. The resistant ones will survive on each side leaving them separated.
4a before the lab:
1. The plasmid may have been supercoiled.
4a Stop and Think:
1. The loading dye is useful because it allows you to see the DNA as it travels through the gel.
2. The DNA may be visible by using a loading dye.
4a Questions:
1. It is important because you want to make sure it will do what you want it to do.
2. My gel results were similar to my predictions.
3. I did not know that there would be a faint band in one of the lanes.
4. The gel does show that we are using the correct plasmid because the base pairs match up.
5. No because there is only one band.
6. In the R+ lane there are 2 bands.
7. You would expect to find the RFP gene in the R+ lane.
8. The DNA ladder did not show up so we were unable to compare the two.
5a before the lab:
1. Growing these ampicillin bacteria could lead to new understandings of medicines.
2. When not given arabinose, these bacteria would die.
3. I predict that the bacteria will grow on both sides of the amp- ara, and only the P+ will grow on the LB-amp.
5a Stop and Think:
1. The purpose of the P- is to be used as a control.
2. The cells need time to recover so that they don't die from the extreme temperatures.
3. Cells are incubated at 37*C because that is the temperature of the human body.
4. You use aseptic techniques so that you won't transfer any bacteria.
5a Questions:
1. Our results matched our predictions.
2. We did not get any red colonies however we did get colonies.
3. The colonies were not resistant to the LB-amp plate.
4. It is important to have multiple copies because you can have extras for cell growth.
5. The RFP gene is expressed as a trait through transcription from DNA to mRNA to protein.
6. Bacteria can again use transcription in the central dogma, and it can become a protein since it is resistant to different viruses, diseases, etc.
6 before the lab:
1. Salt solutions can be used to help the protein stick to the beads and help flush it off the beads.
6 Stop and Think:
1. You can determine where the RFP is because it has a reddish pink color.
2. The supernatant is clear or colorless. The pellet is pink.
3. The BB helps the protein to stick to the beads. The WB gets rid of everything that is not stuck on the beads. The EB flushes the protein.
6 Questions:
1. The eluate containing the RFP gene seemed less bright.
2. The color of the RFP was used as the basis for separation.
3. The precision of the chromatography column could be increased by using the machine which measures everything exactly, or by practicing more.
Analysis/conclusion:
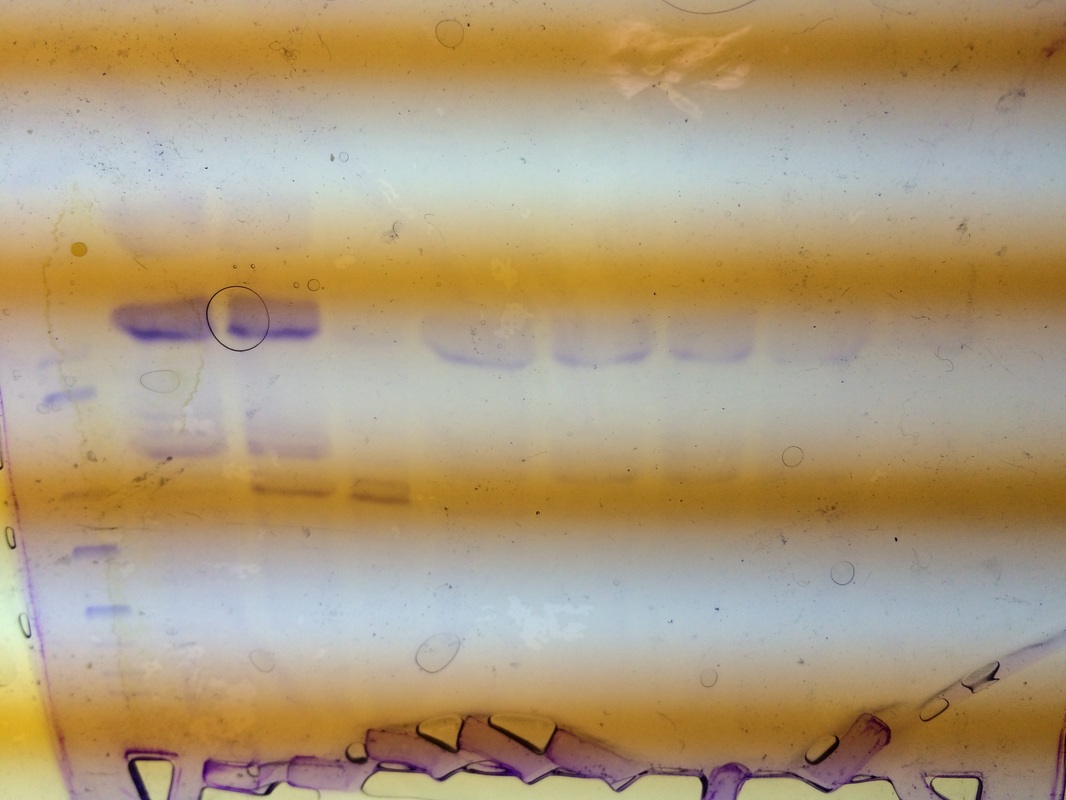
Our RFP was in lane 5. It was pretty light but seemed to be a little below average. It was very pure but not very concentrated. You can tell how concentrated it is by how dark the bands are and you can see how pure it is by the amount of bands in your particular lane.
Reflection:
Our group worked very well together and were able to complete all of the steps correctly. One thing that I would go back and change is trying to get the rest of the members in my group to let me participate a little bit more, I felt as though I didn't get the best experience I could have gotten during this lab because I did not make an effort to really say "can I do this." I just waited for them to assign me something. I only changed this mindset at the end of our lab, and I wish I had done something about it sooner.
Materials and procedure:
Lab 2a can be found in the Amgen lab manual part 2a
Lab 4a can be found in the Amgen lab manual part 2a
Lab 5a can be found in the Amgen lab manual part 5a
Lab 6a can be found in the Amgen lab manual part 6a
Experimental overview:
part 2a: we used restriction digest to verify the plasmid.
-cut plasmid with BamH1 and Hind III to remove RFP-ara from bacterial plasmid
part 4a: we used electrophoresis to verify digest
-to make sure that we had cut the plasmid correctly, we used electrophoresis. This is a process that allows you to run the plasmid in a gel with a DNA ladder.
part 5a: transformation of bacteria with recombinant plasmid
-the biggest part of the lab was in this step. We made the bacteria into the recombinant plasmid. We needed to use enzymes to cut the ligase and plasmid to paste the gene of interest.
part 6: purification of RFP using chromotography
-the last step was to use chromotography to separate the RFP from the rest. We did this by using a column with hydrophilic beads. This allowed the RFP to collect in the resin bed. We then were able to separate the RFP from the beads using a buffer.
Data:
2a before the lab:
1. RFP with pBAD and Ara-C with ori with Amp-R are the two fragments. The RFP and pBAD is 807 Base Pairs, but Ara-C, ori, and Amp-R are 4495 Base Pairs.
2. The components that are needed in the plasmid are Ara-C and the RFP gene.
3. To determine which bacteria will survive you need to use the selectable marker.
2a Stop and Think:
1. The purpose of setting up the tube without the enzymes is to use it as a control sample.
2. The enzymes might work best at 37*C because that is the way it works in the human body so they are comfortable.
2a Questions:
1. Ori = origin of replication; RFP = red fluorescent protein (gene of interest); Amp-R = selectable marker; Ara-C = binds to promoter
2. The restriction enzymes act as a sort of defense weapon.
3. Bacteria would retain certain enzymes to allow them to live. It is making it more difficult in medicine today because the bacteria have become resistant and it is harder to kill them.
4. All follow the same central dogma DNA to mRNA through RNA polymerase, and mRNA to a protein.
5. Put ampicillin and Kanamycin on a petri dish. The resistant ones will survive on each side leaving them separated.
4a before the lab:
1. The plasmid may have been supercoiled.
4a Stop and Think:
1. The loading dye is useful because it allows you to see the DNA as it travels through the gel.
2. The DNA may be visible by using a loading dye.
4a Questions:
1. It is important because you want to make sure it will do what you want it to do.
2. My gel results were similar to my predictions.
3. I did not know that there would be a faint band in one of the lanes.
4. The gel does show that we are using the correct plasmid because the base pairs match up.
5. No because there is only one band.
6. In the R+ lane there are 2 bands.
7. You would expect to find the RFP gene in the R+ lane.
8. The DNA ladder did not show up so we were unable to compare the two.
5a before the lab:
1. Growing these ampicillin bacteria could lead to new understandings of medicines.
2. When not given arabinose, these bacteria would die.
3. I predict that the bacteria will grow on both sides of the amp- ara, and only the P+ will grow on the LB-amp.
5a Stop and Think:
1. The purpose of the P- is to be used as a control.
2. The cells need time to recover so that they don't die from the extreme temperatures.
3. Cells are incubated at 37*C because that is the temperature of the human body.
4. You use aseptic techniques so that you won't transfer any bacteria.
5a Questions:
1. Our results matched our predictions.
2. We did not get any red colonies however we did get colonies.
3. The colonies were not resistant to the LB-amp plate.
4. It is important to have multiple copies because you can have extras for cell growth.
5. The RFP gene is expressed as a trait through transcription from DNA to mRNA to protein.
6. Bacteria can again use transcription in the central dogma, and it can become a protein since it is resistant to different viruses, diseases, etc.
6 before the lab:
1. Salt solutions can be used to help the protein stick to the beads and help flush it off the beads.
6 Stop and Think:
1. You can determine where the RFP is because it has a reddish pink color.
2. The supernatant is clear or colorless. The pellet is pink.
3. The BB helps the protein to stick to the beads. The WB gets rid of everything that is not stuck on the beads. The EB flushes the protein.
6 Questions:
1. The eluate containing the RFP gene seemed less bright.
2. The color of the RFP was used as the basis for separation.
3. The precision of the chromatography column could be increased by using the machine which measures everything exactly, or by practicing more.
Analysis/conclusion:
Our RFP was in lane 5. It was pretty light but seemed to be a little below average. It was very pure but not very concentrated. You can tell how concentrated it is by how dark the bands are and you can see how pure it is by the amount of bands in your particular lane.
Reflection:
Our group worked very well together and were able to complete all of the steps correctly. One thing that I would go back and change is trying to get the rest of the members in my group to let me participate a little bit more, I felt as though I didn't get the best experience I could have gotten during this lab because I did not make an effort to really say "can I do this." I just waited for them to assign me something. I only changed this mindset at the end of our lab, and I wish I had done something about it sooner.